| |
p53 Information
p53 information
p53 story
p53 monoclonal antibodies
p53 pathways
p53 gene
mdm family
mouse models
ASPP family
|
p73 and p63
I am not any more a poor lonesome tumor suppressor gene.
1997 and 1998 have been very prolific with the cloning of two p53 related genes, p73 in 1997 and p63/KET/p51/p40 in 1997/1998.
Kaghad M, Bonnet H, Yang A, Creancier L, Biscan JC, Valent A, Minty A, Chalon P, Lelias JM, Dumont X, Ferrara P, McKeon F, Caput D (1997) Monoallelically expressed gene related to p53 at 1p36, a region frequently deleted in neuroblastoma and other human cancers. Cell 90: 809-819.
Yang AN, Kaghad M, Wang YM, Gillett E, Fleming MD, Dotsch V, Andrews NC, Caput D, McKeon F (1998) p63, a p53 homolog at 3q27-29, encodes multiple products with transactivating, death-inducing, and dominant- negative activities. Mol Cell 2: 305-316.
Trink B, Okami K, Wu L, Sriuranpong V, Jen J, Sidransky D (1998) A new human p53 homologue. Nature Med 4: 747-748.
Schmale H, Bamberger C (1997) A novel protein with strong homology to the tumor suppressor p53. Oncogene 15: 1363-1367.
Lee LA, Walsh P, Prater CA, Su LJ, Marchbank A, Egbert TB, Dellavalle RP, Targoff IN, Kaufman KM, Chorzelski TP, Jablonska S (1999) Characterization of an autoantigen associated with chronic ulcerative stomatitis: the CUSP autoantigen is a member of the p53 family. J Invest Dermatol 113: 146-151. |
p53 gene family
p63 gene organization
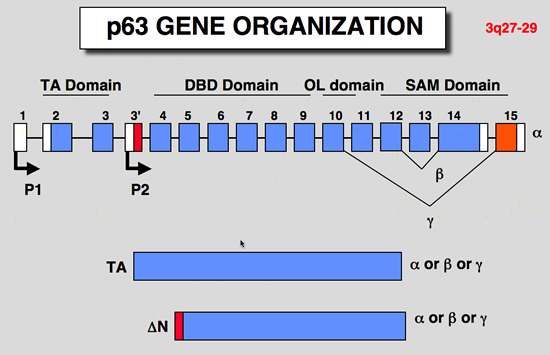 |
| structure of the p63 gene: |
p63 Knock out mice
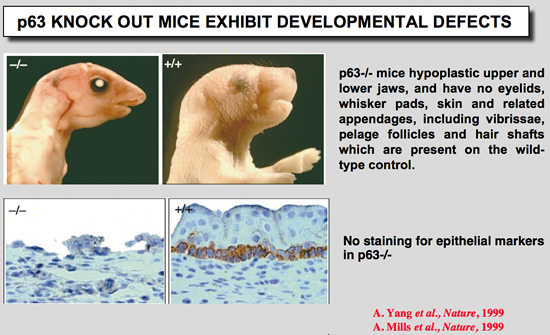 |
Mills, A. A., B. H. Zheng, X. J. Wang, H. Vogel, D. R. Roop, and A. Bradley (1999) Nature 398:708-713.
p63 is a p53 homologue required for limb and epidermal morphogenesis
The p53 tumour suppressor is a transcription factor that regulates the progression of the cell through its cycle and cell death (apoptosis) in response to environmental stimuli such as DNA damage and hypoxia(1,2). Even though p53 modulates these critical cellular processes, mice that lack p53 are developmentally normal(3), suggesting that p53-related proteins might compensate for the functions of p53 during embryogenesis. Two p53 homologues, p63 and p73, are known(4,5) and here we describe the function of p63 in vivo. Mice lacking p63 are born alive but have striking developmental defects. Their limbs are absent or truncated, defects that are caused by a failure of the apical ectodermal ridge to differentiate. The skin of p63- deficient mice does not progress past an early developmental stage: it lacks stratification and does not express differentiation markers. Structures dependent upon epidermal-mesenchymal interactions during embryonic development, such as hair follicles, teeth and mammary glands, are absent in p63-deficient mice. Thus, in contrast to p53, p63 is essential for several aspects of ectodermal differentiation during embryogenesis. |
Yang, A., R. Schweitzer, D. Q. Sun, M. Kaghad, N. Walker, R. T. Bronson, C. Tabin, A. Sharpe, D. Caput, C. Crum, and F. McKeon (1999) Nature 398:714-718.
p63 is essential for regenerative proliferation in limb, craniofacial and epithelial development
The p63 gene, a homologue of the tumour-suppressor p53 (refs 1-5), is highly expressed in the basal or progenitor layers of many epithelial tissues(1). Here we report that mice homozygous for a disrupted p63 gene have major defects in their limb, craniofacial and epithelial development. P63 is expressed in the ectodermal surfaces of the limb buds, branchial arches and epidermal appendages, which are all sites of reciprocal signalling that direct morphogenetic patterning of the underlying mesoderm. The limb truncations are due to a failure to maintain the apical ectodermal ridge, a stratified epithelium, essential for Limb development The embryonic epidermis of p63(-/-) mice undergoes an unusual process of non-regenerative differentiation, culminating in a striking absence of all squamous epithelia and their derivatives, including mammary, lacrymal and salivary glands. Taken together, our results indicate that p63 is critical for maintaining the progenitor-cell populations that are necessary to sustain epithelial development and morphogenesis.
More information on p63 mice models |
p63 and human developmental syndromes
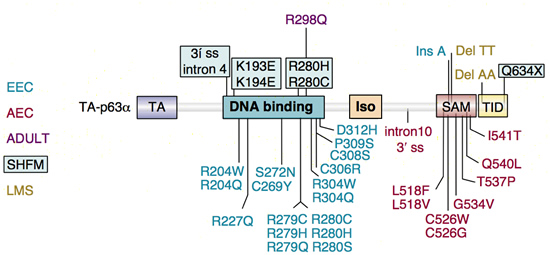 |
p63 mutations in human syndromes. The p63 mutations identified in five different human disorders are indicated on the left.
EEC, ectrodactyly, ectodermal dysplasia, clefting
AEC, ankyloblepharon, ectodermal dysplasia, clefting
ADULT, acro-dermato-ungual-lacrimal-tooth
SHFM, split hand/foot mailformation
LMS, limb-mammary syndrome
Note the clustering of mutations in the DNA-binding domain for ectrodactyly-ectodermal dysplasia-clefting (EEC) syndrome and in the sterile α motif (SAM) domain for ankyloblepharon-ectodermal dysplasia-clefting (AEC) syndrome.
Figure from van Bokhoven, H., and F. McKeon (2002) Trends Mol Med 8:133-139.
Mutations in the p53 homolog p63: allele-specific developmental syndromes in humans
Mutations in p63, the founding member of the p53 family, underlie an array of allele-specific developmental defects involving the limbs and other structures derived from specialized epithelia such as skin, hair, teeth and glands.
p63 structure: transactivation domain (TA), DNA-binding domain and tetramerization domain (Iso). The α isoforms contain in addition a sterile α motif (SAM) domain and a transactivation inhibitory domain (TID) at their C-termini. |
p63 and epithelial stem cell division
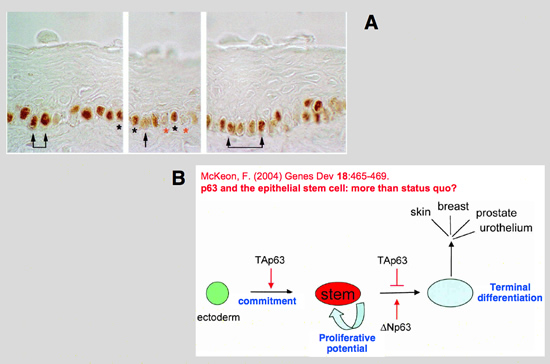 |
| A: In human epidermis, hair follicles, and stratified epidermal cultures, p63 is expressed in the nuclei of cells that are either proliferating or possess the ability to multiply (25).
Staining for p63 revealed that only cells forming the basal layer of human limbus express p63
from: Pellegrini, G., E. Dellambra, O. Golisano, E. Martinelli, I. Fantozzi, S. Bondanza, D. Ponzin, F. McKeon, and M. De Luca (2001) Proc Natl Acad Sci U S A 98:3156-3161.
p63 identifies keratinocyte stem cells.
The proliferative compartment of stratified squamous epithelia consists of stem and transient amplifying (TA) keratinocytes. Some polypeptides are more abundant in putative epidermal stem cells than in TA cells, but no polypeptide confined to the stem cells has yet been identified. Here we show that the p63 transcription factor, a p53 homologue essential for regenerative proliferation in epithelial development, distinguishes human keratinocyte stem cells from their TA progeny. Within the cornea, nuclear p63 is expressed by the basal cells of the limbal epithelium, but not by TA cells covering the corneal surface. Human keratinocyte stem and TA cells when isolated in culture give rise to holoclones and paraclones, respectively. We show by clonal analysis that p63 is abundantly expressed by epidermal and limbal holoclones, but is undetectable in paraclones. TA keratinocytes, immediately after their withdrawal from the stem cell compartment (meroclones), have greatly reduced p63, even though they possess very appreciable proliferative capacity. Clonal evolution (i.e., generation of TA cells from precursor stem cells) is promoted by the sigma isoform of the 14-3-3 family of proteins. Keratinocytes whose 14-3-3final sigma has been down-regulated remain in the stem cell compartment and maintain p63 during serial cultivation. The identification of p63 as a keratinocyte stem cell marker will be of practical importance for the clinical application of epithelial cultures in cell therapy as well as for studies on epithelial tumorigenesis.
B: Assignment of p63 isoform functions in epithelial morphogenesis. Schematic of epithelial morphogenesis depicting the roles ascribed to TAp63 and Np63 isoforms (Koster et al. 2004). In particular, TAp63 is seen to both promote commitment of the ectoderm to epithelial lineages and to suppress the differentiation of such committed progenitor cells. Np63, on the other hand, acts to promote terminal differentiation via its competition and suppression of TAp63 actions in these cells.
from: McKeon, F. (2004) Genes Dev 18:465-469. p63 and the epithelial stem cell: more than status quo? |
p73 gene organisation
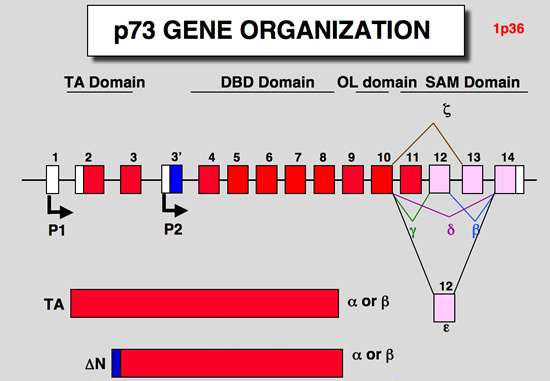 |
| structure of the p73 gene: |
p73 Knock out mice
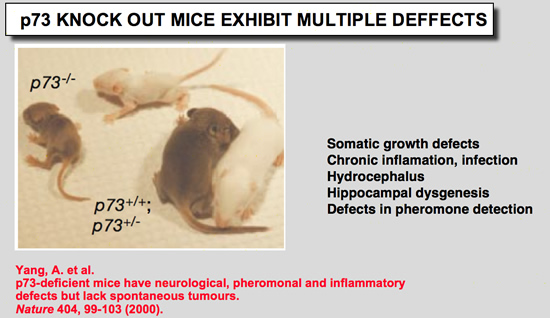 |
Yang, A., N. Walker, R. Bronson, M. Kaghad, M. Oosterwegel, J. Bonnin, C. Vagner, H. Bonnet, P. Dikkes, A. Sharpe, F. McKeon, and D. Caput (2000) Nature 404:99-103.
p73-deficient mice have neurological, pheromonal and inflammatory defects but lack spontaneous tumours
p73 (ref. 1) has high homology with the tumour suppressor p53 (refs 2- 4), as well as with p63, a gene implicated in the maintenance of epithelial stem cells. Despite the localization of the p73 gene to chromosome 1p36.3, a region of frequent aberration in a wide range of human cancers, and the ability of p73 to transactivate p53 target genes, it is unclear whether p73 functions as a tumour suppressor. Here we show that mice functionally deficient for all p73 isoforms exhibit profound defects, including hippocampal dysgenesis, hydrocephalus, chronic infections and inflammation, as well as abnormalities in pheromone sensory pathways. In contrast to p53-deficient mice, however, those lacking p73 show no increased susceptibility to spontaneous tumorigenesis. We report the mechanistic basis of the hippocampal dysgenesis and the loss of pheromone responses, and show that new, potentially dominant-negative, p73 variants are the predominant expression products of this gene in developing and adult tissues. Our data suggest that there is a marked divergence in the physiological functions of the p53 family members, and reveal unique roles for p73 in neurogenesis, sensory pathways and homeostatic control.
More information on p73 mice models |
|
|






